In accordance with SO #3423 - The Gulf of America and SO #3424 - Mount McKinley and Landmarks Honoring the Alaskan People, new USGS data releases specific to those named places will utilize the new name Gulf of America and the restored name Mount McKinley. Per USGS practice, historical data will retain the name of the geographic features as they were known at the time the data were originally released.
Data Release
Cold-Water Coral Microbiomes (Lophelia pertusa) from Gulf of Mexico and Atlantic Ocean: Raw Data
By Christina A. Kellogg and Dawn B. Goldsmith
USGS, St. Petersburg, Florida
Summary
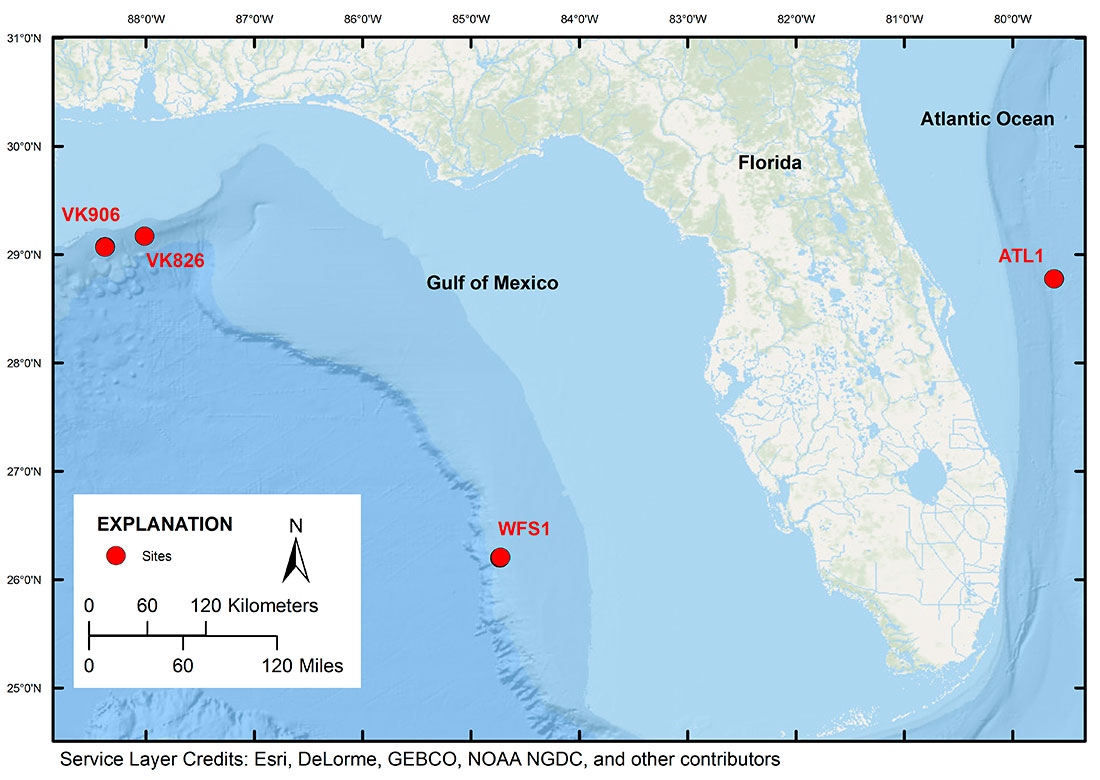
The files in this data release are the raw DNA sequence files referenced in the submitted journal article by Christina A. Kellogg, Dawn B. Goldsmith and Michael A. Gray entitled "Biogeographic comparison of Lophelia-associated bacterial communities in the western Atlantic reveals conserved core microbiome." They represent a 16S rRNA gene amplicon survey of the coral’s microbiomes completed using Roche 454 pyrosequencing with titanium reagents. Samples from the Gulf of Mexico were collected in 2009 and 2010. Samples from the Atlantic Ocean were collected in 2009. The raw data files associated with this study have also been submitted to the NCBI Sequence Read Archive under Bioproject number PRJNA305617. For more information, please see the README file.
Kellogg, C.A., Goldsmith, D.B., and Gray, M.A., 2017, Biogeographic comparison of Lophelia-associated bacterial communities in the western Atlantic reveals conserved core microbiome: Frontiers in Microbiology, 8:796, https://doi.org/10.3389/fmicb.2017.00796.
Data
| File Name and Description | Metadata (XML format) | Metadata (text format) | Download File |
|---|---|---|---|
| Lophelia_raw_data.zip Zip file containing raw data (.fna, .qual, .sff) and MIMARKS metadata. |
Cold_water_coral_microbiomes_Lophelia_pertusa_from_ Gulf_of_Mexico_and_Atlantic_Ocean_raw_data.xml | Cold_water_coral_microbiomes_Lophelia_pertusa_from_ Gulf_of_Mexico_and_Atlantic_Ocean_raw_data.txt |
Lophelia_raw_data.zip (2.6 GB) |
| Supplemental information | |||
| Lophelia_workflow_2016_10_19.txt Details the scripts run in the bioinformatic package QIIME version 1.8 |
Not applicable | Not applicable | Lophelia_workflow_2016_10_19.txt (11 KB) |
| README_Lophelia.txt
Readme file explaining data |
Not applicable | Not applicable | README_Lophelia.txt (5 KB) |
Suggested Citation
Kellogg, C.A., and Goldsmith, D.B., 2017, Cold-water coral microbiomes (Lophelia pertusa) from Gulf of Mexico and Atlantic Ocean: raw data: U.S. Geological Survey data release, https://doi.org/10.5066/F7M32SXM.